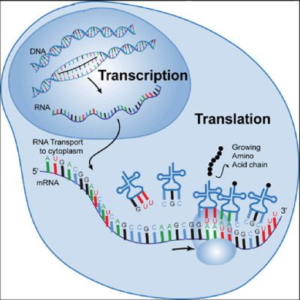
The DNA of a nucleus directs the actions of the cell by instructing it to manufacture specific proteins. This “instruction” is sent from the nucleus into the cytoplasm by “messenger RNA” (mRNA). The mRNA exits the nucleus to bring the “message’” into the cytoplasm, binding with ribosomes to manufacture proteins. Figure 1 illustrates this principle.
In the #5: In Situ Hybridization (ISH) segment of the series, ISH methods were discussed and described as a technique whereby a single strand of nucleotides can be used as a probe to recognize and bind to its complementary strand of DNA – or RNA – located “in place” within the nucleus – or cytoplasm – of cells present in a tissue section.
The current climate in histology favors the development of these new molecular methods and techniques. Why?
More and more treatments for patients are being based upon genetic information from the cell. Pathologists, clinicians and their patients want to obtain as much information as possible from the tissue sections that we prepare in the laboratory. IHC and ISH can localize proteins and nucleic acids within cell nuclei and cytoplasm, as well as in areas outside of the cell. Current methodology allows for double and triple staining of different targets in the same tissue section. This provides extremely valuable information for the pathologist, as single cells can be assessed for the presence or absence of the desired targets within nuclei, cytoplasm and outside of the cell.
A good example of this concept is the current “PIN 4” (prostatic intraepithelial neoplasia) staining procedure for prostate tissue suspected of containing carcinoma cells. Three different proteins are visualized. High molecular weight keratin (HMW) identifies inner luminal cells. P63 protein identifies the basal layer of stratified epithelial cells. These two antibodies are combined in one cocktail and therefore used to identify the basal layers of glandular epithelium within the prostate tissue section. The third protein is identified by an antibody against P504s, which is also called “racemase”. Racemase is an enzyme produced by prostate carcinoma cells.
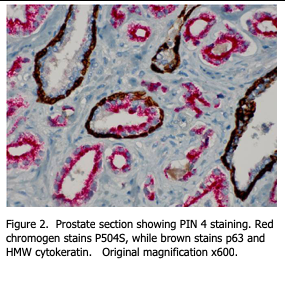
In the double staining method, the HMW / P63 antibody cocktail is applied to the tissue section first. Then a detection chemistry for DAB staining is applied to result in dark brown staining at the antigens’ sites within the basal cell layer. Since both proteins identify the same cell type, it is not necessary to know whether one or both antigens are present – simply that these cells are identified as staining dark brown.
After the stain is completed, a second stain procedure is performed on the same section, using a primary antibody against P504s. For this antibody, a fast red detection chemistry is used such that final stain localization is red. Red staining within cells indicates that the cell is making P506 protein / racemase and is identified as a cancer cell. Figure 2 shows an example of this stain procedure.
This is simply one example of many possibilities using double and triple staining procedures. An excellent paper written by David C. Spaulding describes a specific method for “Simultaneous Detection of Nucleic Acids and Proteins in Pathological Specimens” (1). The paper describes how to identify specific nucleic acids in the nuclei of cells while, in the same section, identifying specific proteins related to these genes.
Clearly, the “molecular revolution” in pathology is underway and histologists should not only be aware of it, but actively participating in it.
References:
- Spaulding D. C. Applications of Combined HIS and IHC Methods for Simultaneous Detection of Nucleic Acids and Proteins in Pathological Specimens. Acta Histochem CytoChem, Vol 28, No 1, 83-84. 1995.
- Chapman C. M. The Histology Handbook. Amazon CreateSpace Independent Publishing Platform; 2017.
- Chapman C. M., Dimenstein I. B. Dermatopathology Laboratory Techniques. Amazon CreateSpace Independent Publishing Platform; 2016.


